import matplotlib.pyplot as plt
import pandas as pd
import numpy as npVolcano Plot (Python)
A volcano plot displays statistical significance (-log10 p-value) versus fold-change (log2 FC) for thousands of features simultaneously. In biomedical research, volcano plots are the standard visualization for differential gene expression results from RNA-seq, proteomics, and metabolomics. Python’s matplotlib provides full control over customizing these publication-ready plots.
Example

Setup
- System Requirements: Cross-platform (Linux/MacOS/Windows)
- Programming Language: Python
- Dependencies:
matplotlib,pandas,numpy
Data Preparation
np.random.seed(42)
n_genes = 5000
df = pd.DataFrame({
'gene': [f'Gene{i+1}' for i in range(n_genes)],
'log2FC': np.random.normal(0, 1.5, n_genes),
'pvalue': np.random.uniform(1e-10, 1, n_genes)
})
df['neg_log10p'] = -np.log10(df['pvalue'])
fc_thresh = 1.0
p_thresh = 0.05
conditions = [
(df['log2FC'] > fc_thresh) & (df['pvalue'] < p_thresh),
(df['log2FC'] < -fc_thresh) & (df['pvalue'] < p_thresh),
]
choices = ['Up', 'Down']
df['regulation'] = np.select(conditions, choices, default='NS')Visualization
Basic Volcano Plot
colors = {'Up': '#e63946', 'Down': '#457b9d', 'NS': '#cccccc'}
fig, ax = plt.subplots(figsize=(8, 6))
for reg, color in colors.items():
subset = df[df['regulation'] == reg]
ax.scatter(subset['log2FC'], subset['neg_log10p'],
c=color, s=8, alpha=0.6, label=f'{reg} ({len(subset)})')
ax.axhline(-np.log10(p_thresh), color='grey', linestyle='--', linewidth=0.8)
ax.axvline(fc_thresh, color='grey', linestyle='--', linewidth=0.8)
ax.axvline(-fc_thresh, color='grey', linestyle='--', linewidth=0.8)
ax.set_xlabel('log₂(Fold Change)')
ax.set_ylabel('-log₁₀(P-value)')
ax.set_title('Volcano Plot')
ax.legend(frameon=False)
ax.spines[['top', 'right']].set_visible(False)
plt.tight_layout()
plt.show()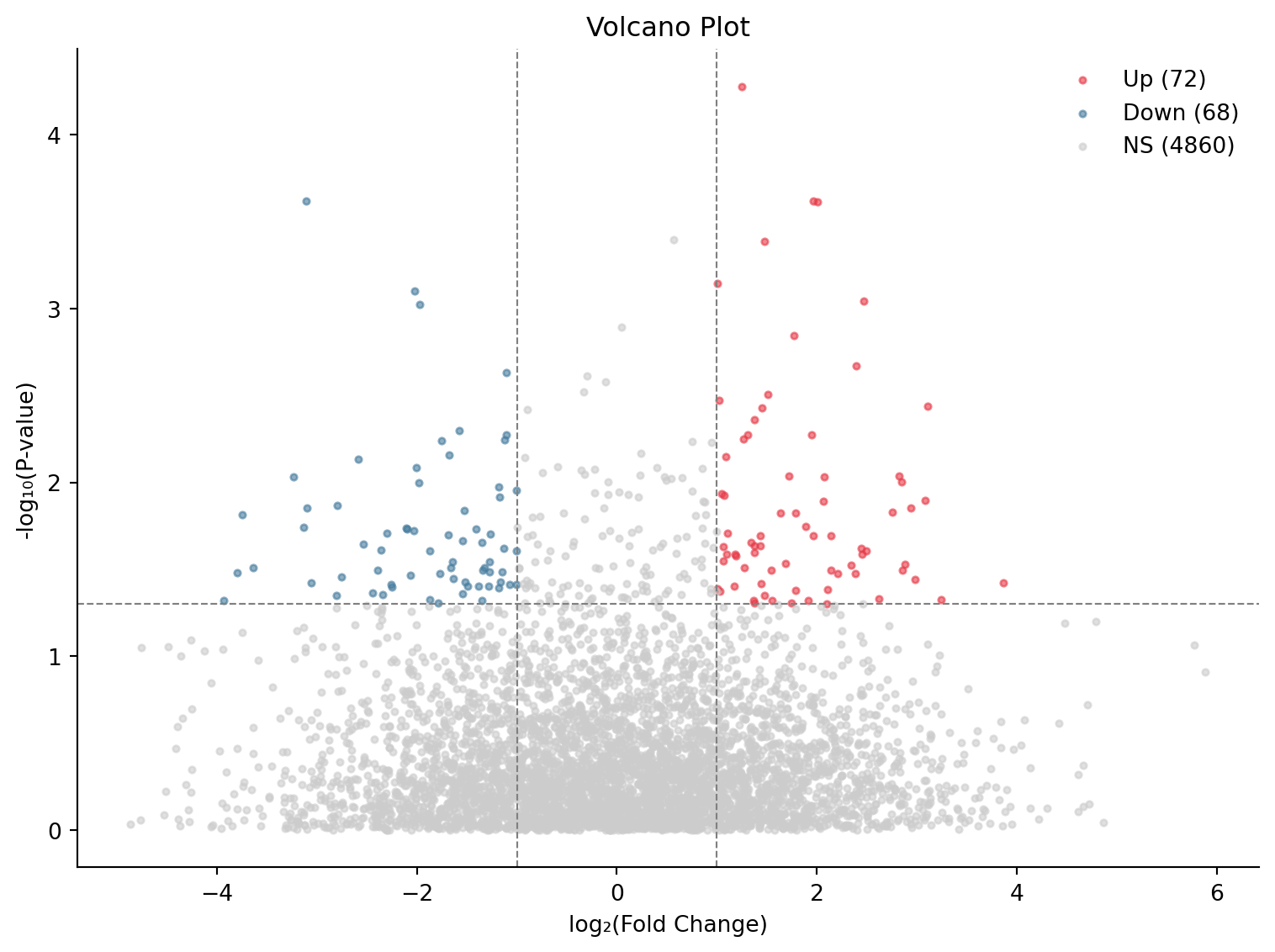
Volcano Plot with Gene Labels
fig, ax = plt.subplots(figsize=(9, 7))
for reg, color in colors.items():
subset = df[df['regulation'] == reg]
ax.scatter(subset['log2FC'], subset['neg_log10p'],
c=color, s=8, alpha=0.5, label=f'{reg} ({len(subset)})')
top_genes = df.nlargest(15, 'neg_log10p')
for _, row in top_genes.iterrows():
ax.annotate(row['gene'], (row['log2FC'], row['neg_log10p']),
fontsize=7, ha='center', va='bottom',
arrowprops=dict(arrowstyle='-', color='grey', lw=0.5))
ax.axhline(-np.log10(p_thresh), color='grey', linestyle='--', linewidth=0.8)
ax.axvline(fc_thresh, color='grey', linestyle='--', linewidth=0.8)
ax.axvline(-fc_thresh, color='grey', linestyle='--', linewidth=0.8)
ax.set_xlabel('log₂(Fold Change)')
ax.set_ylabel('-log₁₀(P-value)')
ax.set_title('Volcano Plot with Top Gene Labels')
ax.legend(frameon=False, loc='upper left')
ax.spines[['top', 'right']].set_visible(False)
plt.tight_layout()
plt.show()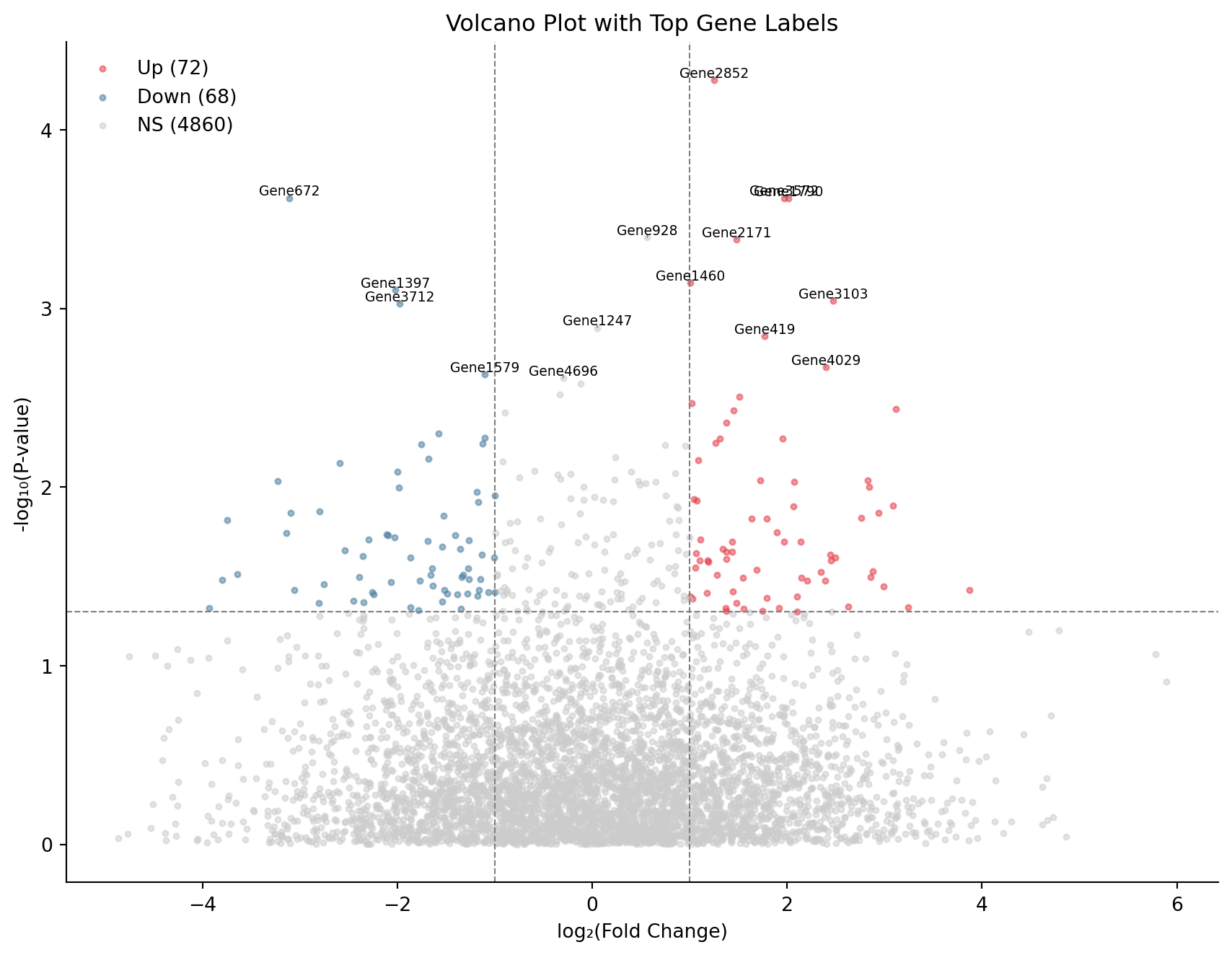
Enhanced Volcano with Significance Regions
fig, ax = plt.subplots(figsize=(9, 7))
sig_up = df[(df['regulation'] == 'Up')]
sig_down = df[(df['regulation'] == 'Down')]
ns = df[df['regulation'] == 'NS']
ax.scatter(ns['log2FC'], ns['neg_log10p'], c='#e0e0e0', s=6, alpha=0.4, label=f'NS ({len(ns)})')
ax.scatter(sig_down['log2FC'], sig_down['neg_log10p'], c='#457b9d', s=12, alpha=0.7, label=f'Down ({len(sig_down)})')
ax.scatter(sig_up['log2FC'], sig_up['neg_log10p'], c='#e63946', s=12, alpha=0.7, label=f'Up ({len(sig_up)})')
ax.axhline(-np.log10(p_thresh), color='black', linestyle=':', linewidth=0.8, alpha=0.5)
ax.axvline(fc_thresh, color='black', linestyle=':', linewidth=0.8, alpha=0.5)
ax.axvline(-fc_thresh, color='black', linestyle=':', linewidth=0.8, alpha=0.5)
ax.fill_betweenx([0, df['neg_log10p'].max()], fc_thresh, df['log2FC'].max(),
alpha=0.03, color='red')
ax.fill_betweenx([0, df['neg_log10p'].max()], df['log2FC'].min(), -fc_thresh,
alpha=0.03, color='blue')
ax.set_xlabel('log₂(Fold Change)', fontsize=12)
ax.set_ylabel('-log₁₀(P-value)', fontsize=12)
ax.set_title('Enhanced Volcano Plot', fontsize=14)
ax.legend(frameon=True, fancybox=True, shadow=True)
ax.spines[['top', 'right']].set_visible(False)
plt.tight_layout()
plt.show()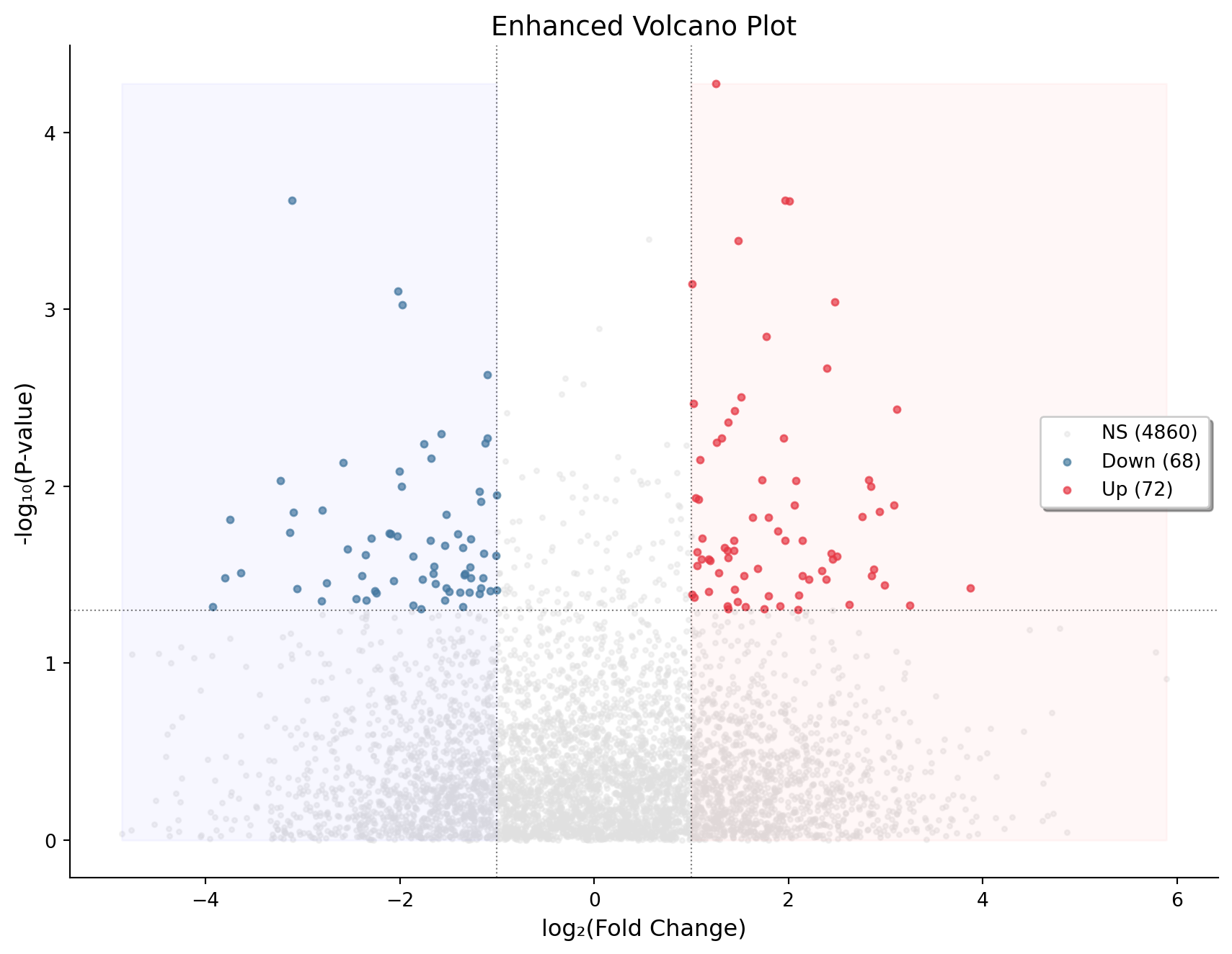
References
- Li, W. (2012). Volcano plots in analyzing differential expressions with mRNA microarrays. Journal of Bioinformatics and Computational Biology, 10(6), 1231003.