# Install Bioconductor manager if needed
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
# Install CRAN dependencies
if (!requireNamespace("ggplot2", quietly = TRUE)) install.packages("ggplot2")
if (!requireNamespace("circlize", quietly = TRUE)) install.packages("circlize")
if (!requireNamespace("igraph", quietly = TRUE)) install.packages("igraph")
# Install CellChat from GitHub (jinworks/CellChat is the current maintained fork)
if (!requireNamespace("CellChat", quietly = TRUE)) {
if (!requireNamespace("remotes", quietly = TRUE)) install.packages("remotes")
remotes::install_github("jinworks/CellChat")
}
# Load packages
library(CellChat)
library(ggplot2)
library(circlize)Cell-Cell Communication Circle Plot
The Cell-Cell Communication Circle Plot (细胞-细胞通讯网络圈图) is a specialized visualization for depicting intercellular signaling interactions inferred from single-cell RNA sequencing (scRNA-seq) data. Using the CellChat R package, this plot presents a circular network where nodes represent cell populations (cell types or clusters) and directed edges indicate the strength and direction of ligand-receptor communication signals between them.
The arc width of each edge encodes communication probability, and the self-loops on each node represent autocrine signaling. Color coding maps each cell type to a distinct hue, and arrow directionality shows the sender-receiver relationship. This visualization is particularly powerful for discovering cross-talk between immune cells and tumor cells, identifying key signaling hubs in tissue microenvironments, and comparing communication networks across biological conditions.
Example
Setup
System Requirements: Cross-platform (Linux/MacOS/Windows)
Programming Language: R
Dependencies:
CellChat,ggplot2,circlize
sessioninfo::session_info("attached")─ Session info ───────────────────────────────────────────────────────────────
setting value
version R version 4.6.0 (2026-04-24)
os Ubuntu 24.04.4 LTS
system x86_64, linux-gnu
ui X11
language (EN)
collate C.UTF-8
ctype C.UTF-8
tz UTC
date 2026-05-05
pandoc 3.1.3 @ /usr/bin/ (via rmarkdown)
quarto 1.9.37 @ /usr/local/bin/quarto
─ Packages ───────────────────────────────────────────────────────────────────
package * version date (UTC) lib source
Biobase * 2.72.0 2026-04-28 [1] Bioconduc~
BiocGenerics * 0.58.0 2026-04-28 [1] Bioconduc~
CellChat * 2.2.0.9001 2026-05-04 [1] Github (jinworks/CellChat@75253cd)
circlize * 0.4.18 2026-04-04 [1] RSPM
dplyr * 1.2.1 2026-04-03 [1] RSPM
generics * 0.1.4 2025-05-09 [1] RSPM
ggplot2 * 4.0.3.9000 2026-05-04 [1] Github (tidyverse/ggplot2@6870419)
igraph * 2.3.0 2026-04-21 [1] RSPM
[1] /home/runner/work/_temp/Library
[2] /opt/R/4.6.0/lib/R/site-library
[3] /opt/R/4.6.0/lib/R/library
* ── Packages attached to the search path.
──────────────────────────────────────────────────────────────────────────────Data Preparation
For this tutorial, we build a simulated CellChat-compatible communication matrix so the example is fully self-contained and runs without requiring an external dataset or model inference. The same visualizations work identically with a real CellChat object produced from your own scRNA-seq data.
# Simulate a communication count/weight matrix
# In real use, these come from cellchat@net$count and cellchat@net$weight
set.seed(42)
cell_types <- c("CD4 T", "CD8 T", "NK", "B cell", "Monocyte",
"DC", "Fibroblast", "Epithelial", "Endothelial")
n <- length(cell_types)
net_count <- matrix(sample(0:50, n * n, replace = TRUE), n, n,
dimnames = list(cell_types, cell_types))
net_weight <- matrix(runif(n * n, 0, 1), n, n,
dimnames = list(cell_types, cell_types))
diag(net_count) <- 0 # remove self-communication
diag(net_weight) <- 0
# Group size (proportional to number of cells per type, simulated)
group_size <- sample(50:500, n, replace = TRUE)
names(group_size) <- cell_types
cat("Simulated", n, "cell types with", sum(net_count), "total interactions\n")Simulated 9 cell types with 1875 total interactionsprint(net_count) CD4 T CD8 T NK B cell Monocyte DC Fibroblast Epithelial Endothelial
CD4 T 0 23 46 4 29 44 48 37 41
CD8 T 36 0 2 19 42 27 25 9 8
NK 0 35 0 33 14 4 49 39 28
B cell 24 24 24 0 21 3 5 4 11
Monocyte 9 36 26 39 0 33 5 32 19
DC 35 45 35 2 35 0 1 48 8
Fibroblast 17 19 36 32 3 34 0 38 42
Epithelial 48 25 30 41 21 23 20 0 34
Endothelial 46 49 44 23 17 22 1 44 0Build a CellChat Object from Your Own Data (Reference)
The following code is for reference only (eval: false). It shows how to build a CellChat object from a Seurat scRNA-seq object before running the visualizations below.
library(Seurat)
library(CellChat)
# 1. Extract normalised count matrix and metadata from a Seurat object
seurat_obj <- readRDS("your_seurat_object.rds")
data_input <- GetAssayData(seurat_obj, assay = "RNA", slot = "data")
meta <- seurat_obj@meta.data
# 2. Create CellChat object
cellchat <- createCellChat(object = data_input, meta = meta, group.by = "cell_type")
# 3. Set ligand-receptor database
cellchat@DB <- CellChatDB.human # or CellChatDB.mouse
# 4. Pre-process & infer communication
cellchat <- subsetData(cellchat)
cellchat <- identifyOverExpressedGenes(cellchat)
cellchat <- identifyOverExpressedInteractions(cellchat)
cellchat <- computeCommunProb(cellchat, type = "triMean")
cellchat <- filterCommunication(cellchat, min.cells = 10)
cellchat <- computeCommunProbPathway(cellchat)
cellchat <- aggregateNet(cellchat)
# Now cellchat@net$count and cellchat@net$weight are available for plottingVisualization
1. Circle Plot of Overall Cell-Cell Communication Network
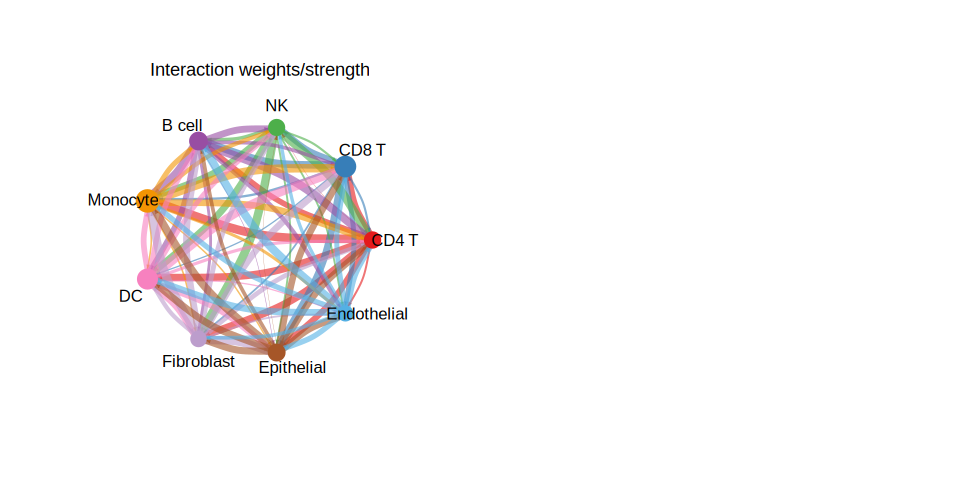
The overall circle plot summarizes all inferred communication interactions. Each node represents a cell type; node size is proportional to the number of interactions it participates in. Arc width encodes communication strength.
par(mfrow = c(1, 2), xpd = TRUE)
# Panel A: Number of interactions
netVisual_circle(
net_count,
vertex.weight = group_size,
weight.scale = TRUE,
label.edge = FALSE,
title.name = "Number of interactions"
)
# Panel B: Interaction strength (weights)
netVisual_circle(
net_weight,
vertex.weight = group_size,
weight.scale = TRUE,
label.edge = FALSE,
title.name = "Interaction weights/strength"
)
par(mfrow = c(1, 1))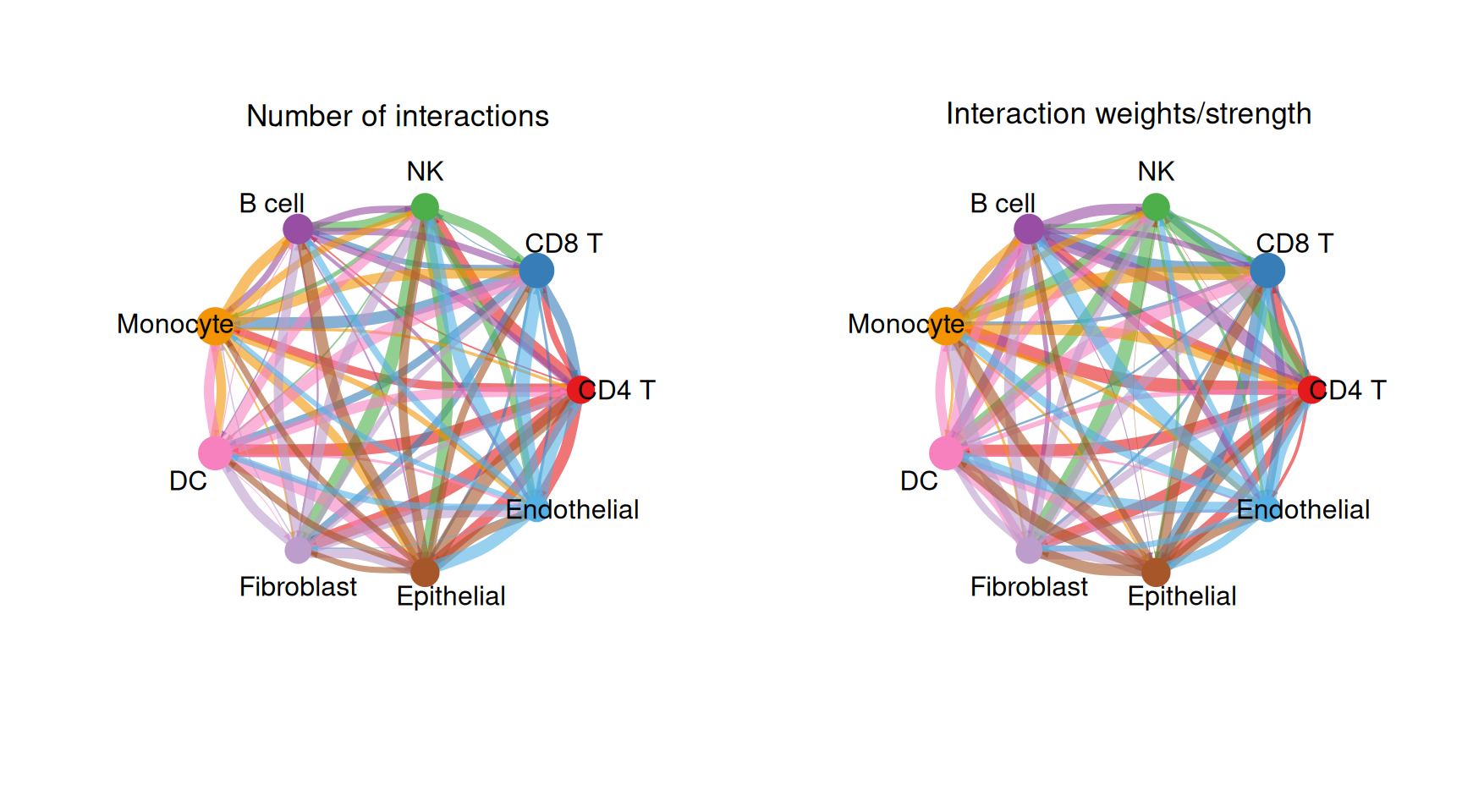
2. Circle Plot for Individual Cell Groups
To highlight the outgoing signals from each cell type separately, we subset the weight matrix row by row and produce one circle per cell type.
n_types <- nrow(net_weight)
n_cols <- ceiling(sqrt(n_types))
n_rows <- ceiling(n_types / n_cols)
par(mfrow = c(n_rows, n_cols), xpd = TRUE, mar = c(1, 1, 2, 1))
for (i in seq_len(n_types)) {
mat_i <- matrix(0, nrow = n_types, ncol = n_types,
dimnames = dimnames(net_weight))
mat_i[i, ] <- net_weight[i, ] # outgoing signals from cell type i
netVisual_circle(
mat_i,
vertex.weight = group_size,
weight.scale = TRUE,
edge.weight.max = max(net_weight),
title.name = rownames(net_weight)[i]
)
}
par(mfrow = c(1, 1))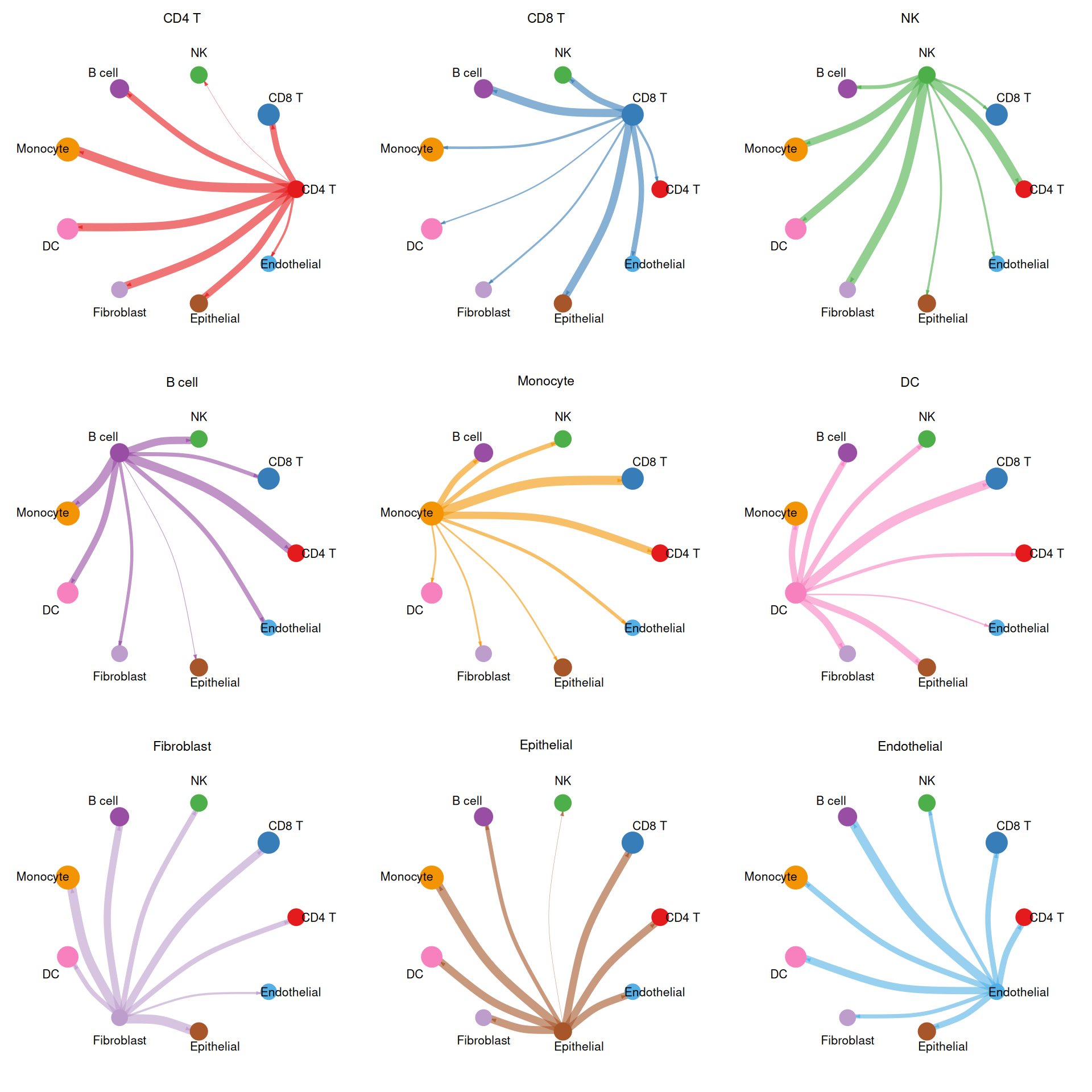
3. Chord Diagram
The chord diagram provides an alternative circular view using circlize, clearly showing bidirectional communication volumes between cell type pairs.
# Use circlize for a chord diagram of communication weights
chordDiagram(
net_weight,
transparency = 0.4,
annotationTrack = "grid",
preAllocateTracks = 1
)
circos.trackPlotRegion(
track.index = 1,
panel.fun = function(x, y) {
circos.text(
CELL_META$xcenter,
CELL_META$ylim[1] + 0.1,
CELL_META$sector.index,
facing = "clockwise",
niceFacing = TRUE,
adj = c(0, 0.5),
cex = 0.7
)
},
bg.border = NA
)
title("Cell-Cell Communication (Chord Diagram)")
circos.clear()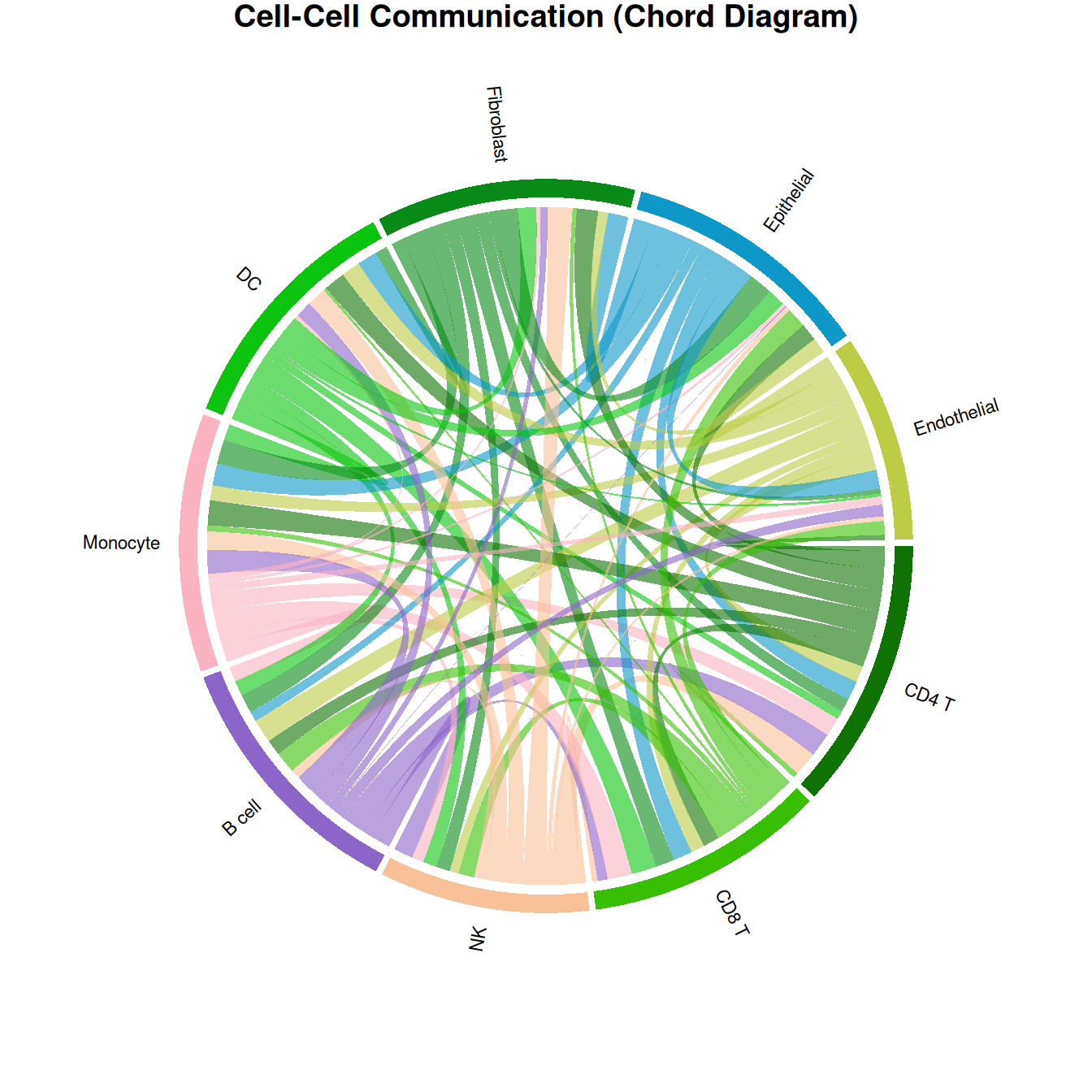
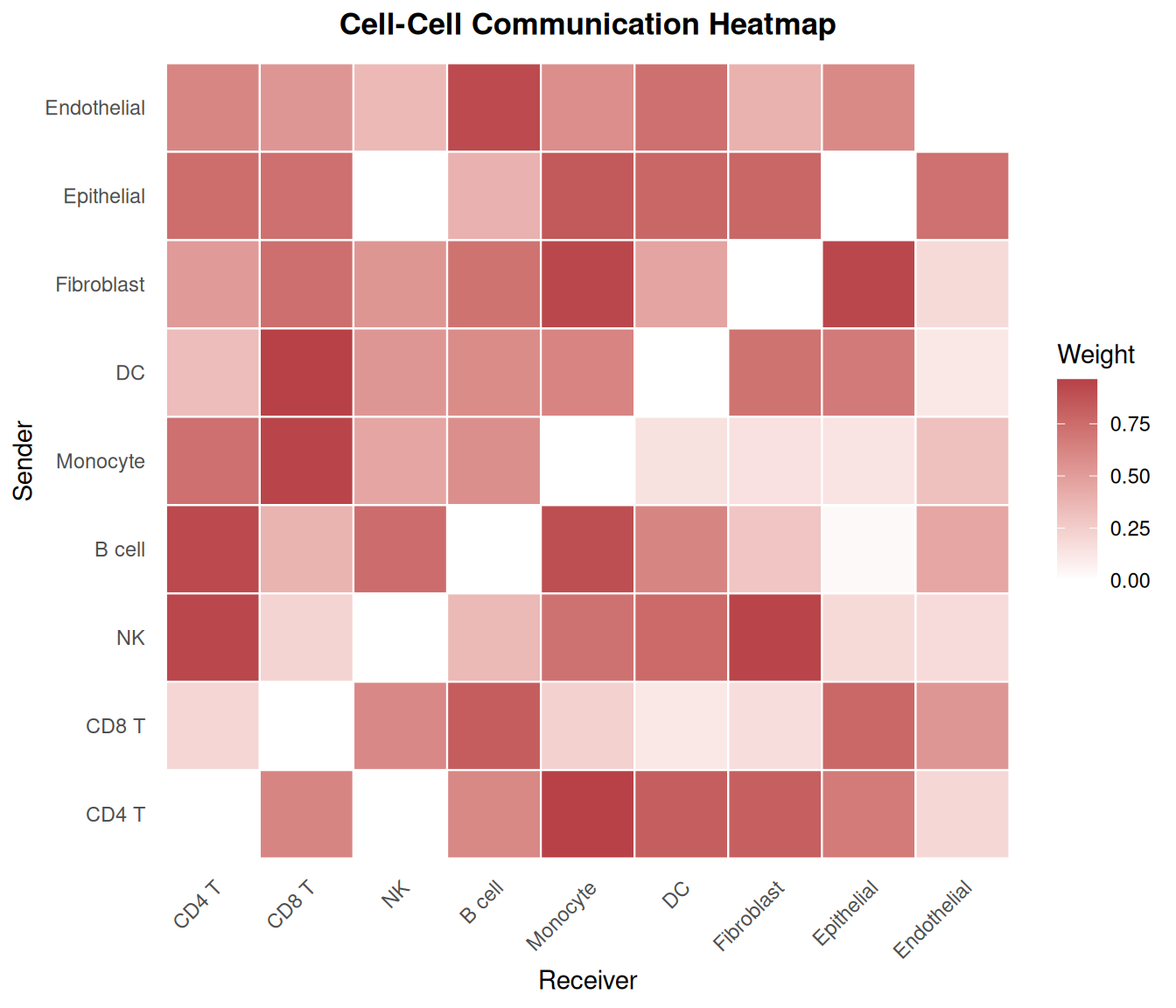
4. Communication Heatmap
A heatmap view allows rapid identification of dominant sender-receiver pairs and asymmetric communication patterns.
df_heat <- as.data.frame(as.table(net_weight))
colnames(df_heat) <- c("Sender", "Receiver", "Weight")
ggplot(df_heat, aes(x = Receiver, y = Sender, fill = Weight)) +
geom_tile(color = "white", linewidth = 0.4) +
scale_fill_gradient(low = "white", high = "#b74147", name = "Weight") +
theme_minimal(base_size = 11) +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
panel.grid = element_blank(),
plot.title = element_text(size = 13, face = "bold", hjust = 0.5)
) +
labs(
title = "Cell-Cell Communication Heatmap",
x = "Receiver", y = "Sender"
)
References
[1] Jin S, Guerrero-Juarez CF, Zhang L, et al. Inference and analysis of cell-cell communication using CellChat. Nature Communications. 2021;12(1):1088. https://doi.org/10.1038/s41467-021-21246-9
[2] Jin S, et al. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nature Protocols. 2024. https://doi.org/10.1038/s41596-024-01045-4
[3] CellChat GitHub: https://github.com/jinworks/CellChat